| y | cg11012046 | cg21859434 | cg23970089 | cg25176823 | cg24576735 | age | cg19099850 | cg17403875 | cg24078828 |
|---|---|---|---|---|---|---|---|---|---|
| Mesenchymal-like | 0.07659845 | 0.013786153 | 0.3651860 | 0.32445298 | 0.03255164 | 44 | 0.04020459 | 0.01333304 | 0.02021659 |
| Mesenchymal-like | 0.15108266 | 0.009155272 | 0.6024613 | 0.36269407 | 0.02215452 | 50 | 0.03771707 | 0.01045362 | 0.02098629 |
| Classic-like | 0.42735137 | 0.380247358 | 0.1984322 | 0.27367605 | 0.04060163 | 61 | 0.03648317 | 0.01304019 | 0.02570674 |
| G-CIMP-low | 0.93509040 | 0.861978174 | 0.5049500 | 0.47706717 | 0.03564740 | 20 | 0.88262115 | 0.62531752 | 0.42465204 |
| LGm6-GBM | 0.04602123 | 0.804717273 | 0.2564513 | 0.03827349 | 0.03625206 | 18 | 0.82269945 | 0.80449412 | 0.78443442 |
Tree-Based Machine Learning Methods for Prediction and Variable Selection
Part II
University of Miami
Inference
| Family | Terminal Node Statistics, Prediction |
|---|---|
| Regression Quantile Regression |
mean, mean quantiles, moments, mean |
| Classification Imbalanced Two-Class |
class proportions, class proportions, Bayes classifier class proportions, class proportions, q-classifier |
| Survival | Kaplan-Meier survival, Nelson-Aalen CHF, mortality |
| Competing Risk | cause-specific CHF, cause-specific CIF, event-specific expected number of years lost |
| Multivariate Regression Multivariate Classification \(\quad\) Multivariate Mixed Regression Multivariate Quantile Regression |
per response: mean, mean per response: class proportions, class proportions, Bayes classifier same as above for Regression, Classification per response: quantiles, mean |
| Unsupervised sidClustering Breiman (Shi-Horvath) |
none same as Multivariate Mixed Regression same as Classification |
Inference
RF actually produces two ensembles!
The inbag ensemble is the averaged ensemble over the trained trees, where each tree is trained by a subsampled (bootstrap data) \[ F_{\!\text{ens}}(\mathbf{x}) =\frac{1}{B}\sum_{b=1}^B h^*_b(\mathbf{x}) \]
The out-of-bag (OOB) ensemble is case-specific. For case \(i\), the average is taken over trees where \(i\) is OOB \[ F_{\!\text{ens}}^{(i)} =\frac{1}{\#O_i}\sum_{b\in O_i} h^*_b(\mathbf{x}_i),\hskip10pt \#O_i\approx .368 B \]
The inbag ensemble is used for prediction on test data. The OOB ensemble is used for inference
OOB ensemble
- Equivalent to a leave-one-out (loo) bootstrap estimator [1]. Therefore it can be used for estimating generalization error \[ \text{Err} = \frac{1}{n}\sum_{i=1}^n L(Y_i, F_{\!\text{ens}}^{(i)}) \] For example, for regression \[ \text{Prediction Error} = \frac{1}{n}\sum_{i=1}^n (Y_i-F_{\!\text{ens}}^{(i)})^2 \]
- Avoids the need for cross-validation. Very conveniently for prediction error and for tuning the forest.
- Provides cross-validated estimator for model – a unique feature not seen with other ML methods
Key quantities for classification
The key quantities returned by the package are
OOB classification example: Glioma
o <- rfsrc(y~., data = glioma)
o
Sample size: 878
Frequency of class labels: Classic-like=148, Codel=174, G-CIMP-high=249, G-CIMP-low=25, LGm6-GBM=41, Mesenchymal-like=215, PA-like=26
Number of trees: 500
Forest terminal node size: 1
Average no. of terminal nodes: 73.896
No. of variables tried at each split: 36
Total no. of variables: 1241
Resampling used to grow trees: swor
Resample size used to grow trees: 555
Analysis: RF-C
Family: class
Splitting rule: gini *random*
Number of random split points: 10
(OOB) Brier score: 0.02292715
(OOB) Normalized Brier score: 0.18723839
(OOB) AUC: 0.99338879
(OOB) Log-loss: 0.33872539
(OOB) Requested performance error: 0.07061503, 0.05405405, 0.01149425, 0.00803213, 0.52, 0.51219512, 0.06046512, 0.11538462
Confusion matrix:
predicted
observed Classic-like Codel G-CIMP-high G-CIMP-low LGm6-GBM Mesenchymal-like PA-like class.error
Classic-like 140 0 0 0 0 8 0 0.0541
Codel 0 172 2 0 0 0 0 0.0115
G-CIMP-high 0 2 247 0 0 0 0 0.0080
G-CIMP-low 0 0 13 12 0 0 0 0.5200
LGm6-GBM 0 1 0 0 20 20 0 0.5122
Mesenchymal-like 13 0 0 0 0 202 0 0.0605
PA-like 0 0 0 0 0 3 23 0.1154
(OOB) Misclassification rate: 0.07061503
Random-classifier baselines (uniform):
Brier: 0.12244898 Normalized Brier: 1 Log-loss: 1.94591015
mean(o$class!=o$yvar)
> [1] 0
# inbag misclassification is zero!
mean(o$class.oob!=o$yvar)
> [1] 0.07061503:::
Key quantities
The key quantities returned by the package are
o$predicted ---> inbag estimated probabilities
o$predicted.oob ---> OOB estimated probabilities
o$class ---> inbag class predictions
o$class.oob ---> OOB class predictionso$predicted[1:5,] # ---> inbag estimated probabilities
> Classic-like Codel G-CIMP-high G-CIMP-low LGm6-GBM Mesenchymal-like PA-like
> [1,] 0.006 0.002 0.000 0.004 0.084 0.892 0.012
> [2,] 0.136 0.008 0.000 0.004 0.014 0.832 0.006
> [3,] 0.064 0.004 0.008 0.010 0.018 0.892 0.004
> [4,] 0.928 0.008 0.004 0.004 0.002 0.054 0.000
> [5,] 0.936 0.000 0.000 0.002 0.000 0.062 0.000
o$predicted.oob[1:5,] # ---> OOB estimated probabilities
> Classic-like Codel G-CIMP-high G-CIMP-low LGm6-GBM Mesenchymal-like PA-like
> [1,] 0.01840491 0.006134969 0.00000000 0.012269939 0.257668712 0.6687117 0.03680982
> [2,] 0.38857143 0.022857143 0.00000000 0.011428571 0.040000000 0.5200000 0.01714286
> [3,] 0.17391304 0.010869565 0.02173913 0.027173913 0.048913043 0.7065217 0.01086957
> [4,] 0.81443299 0.020618557 0.01030928 0.010309278 0.005154639 0.1391753 0.00000000
> [5,] 0.82222222 0.000000000 0.00000000 0.005555556 0.000000000 0.1722222 0.00000000
o$class[1:5] # ---> inbag class predictions
> [1] Mesenchymal-like Mesenchymal-like Mesenchymal-like Classic-like Classic-like
o$class.oob[1:5] # ---> OOB class predictions
> [1] Mesenchymal-like Mesenchymal-like Mesenchymal-like Classic-like Classic-like Key quantities for survival
The key quantities returned by the package are
o$time.interest ---> event times (everything keys off this)
o$predicted ---> inbag estimated mortality
o$predicted.oob ---> OOB estimated mortality
o$survival ---> inbag survival estimator for each case
o$survival.oob ---> OOB survival estimator for each case
o$chf ---> inbag CHF estimator for each case
o$chf.oob ---> OOB CHF estimator for each caseOOB survival example: PBC Mayo Clinic
## load the PBC data
data(pbc, package="survival")
## remove the ID
pbc$id <- NULL
## convert to right-censoring with death as the event
pbc$status[pbc$status > 0] <- 1
## default RSF call
o <- rfsrc(Surv(time, status)~., pbc)
## choose some cases
idx <- c(11,34,60)
## plot the curves
matplot(o$time.interest,
t(o$survival[idx,]), type = "l", col=4, lwd=3,
xlab = "Days", ylab = "Survival")
matlines(o$time.interest,
t(o$survival.oob[idx,]), type = "l", col=2, lwd=3)
legend("bottomleft", legend = c("inbag", "oob"), fill = c(4,2))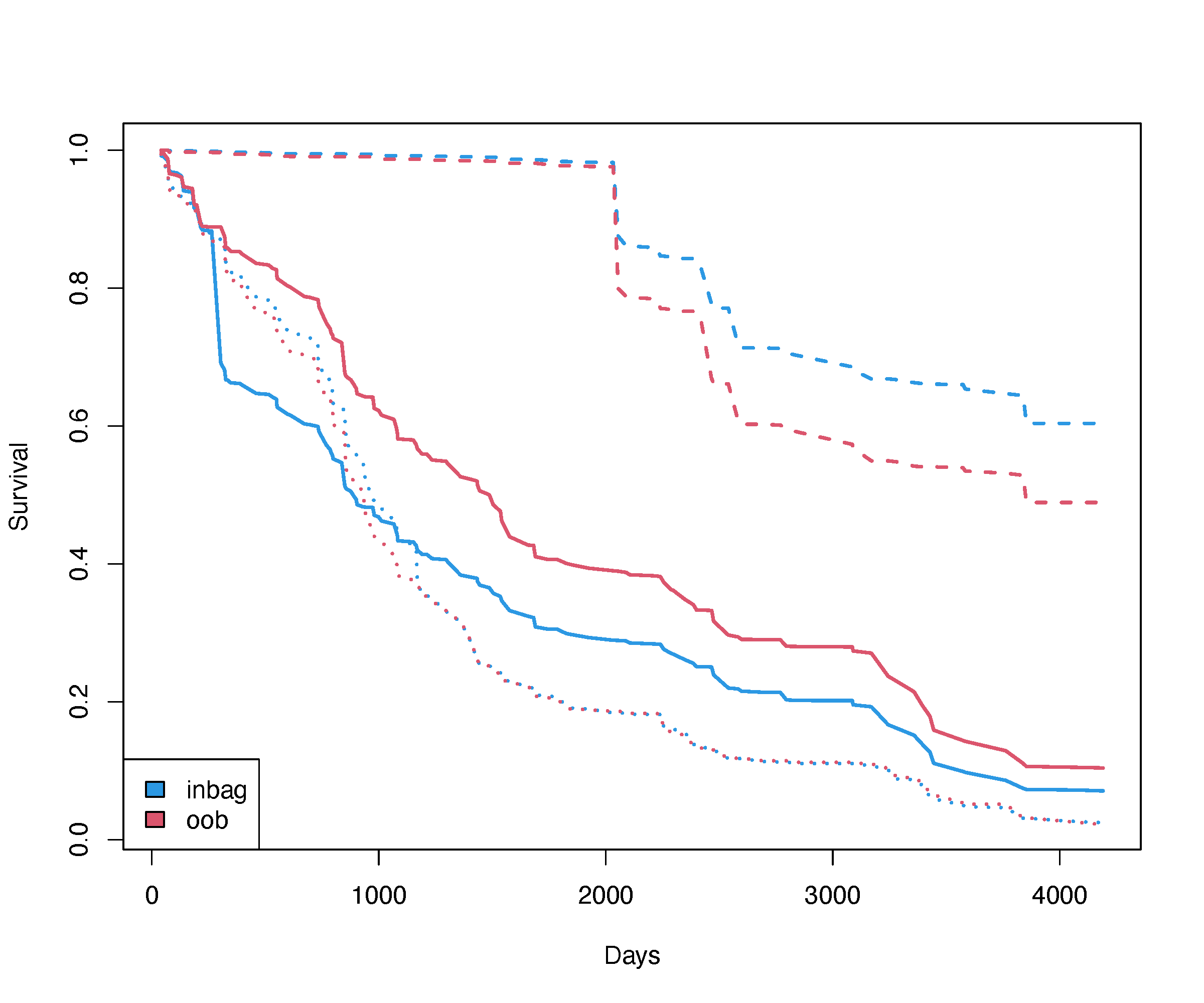
Key quantities
Prediction error
| Family | Prediction Error |
|---|---|
| Regression Quantile Regression |
mean-squared error mean-squared error |
| Classification Imbalanced Two-Class |
misclassification, Brier score G-mean, misclassification, Brier score |
| Survival | Harrell’s C-index (1 minus concordance) |
| Competing Risk | Harrell’s C-index (1 minus concordance) |
| Multivariate Regression Multivariate Classification Multivariate Mixed Regression Multivariate Quantile Regression |
per response: same as above for Regression per response: same as above for Classification per response: same as above for Regression, Classification same as Multivariate Regression |
| Unsupervised sidClustering Breiman (Shi-Horvath) |
none same as Multivariate Mixed Regression same as Classification |
Prediction error for classification
Error rate for the chosen performance metric is always available in obj$err.rate, however any error rate can be calculated using obj$yvar with obj$predicted.oob or obj$class.oob
- Misclassification error:
get.misclass.error - Brier error:
get.brier.error - Log-loss (cross-entropy loss):
get.logloss - AUC:
get.auc
Classification example: Glioma
Different prediction measures
Prediction error for survival
1. Harrell’s C-error rate
Extract using
obj$err.rateorget.cindexEasy to understand
Computationally expensive
Biased in the presence of heavy censoring
2. Time-varying Brier score
- Not intuitive
- Computationally inexpensive:
get.brier.survival - Unbiased and robust
3. Time-varying AUC (AUC-t)
- Easy to understand
- In principle robust
- Requires sophisticated methods for calculating (see
riskRegressionpackage)
Prediction error for survival
Time-varying brier score and AUC-t for the PBC dataset
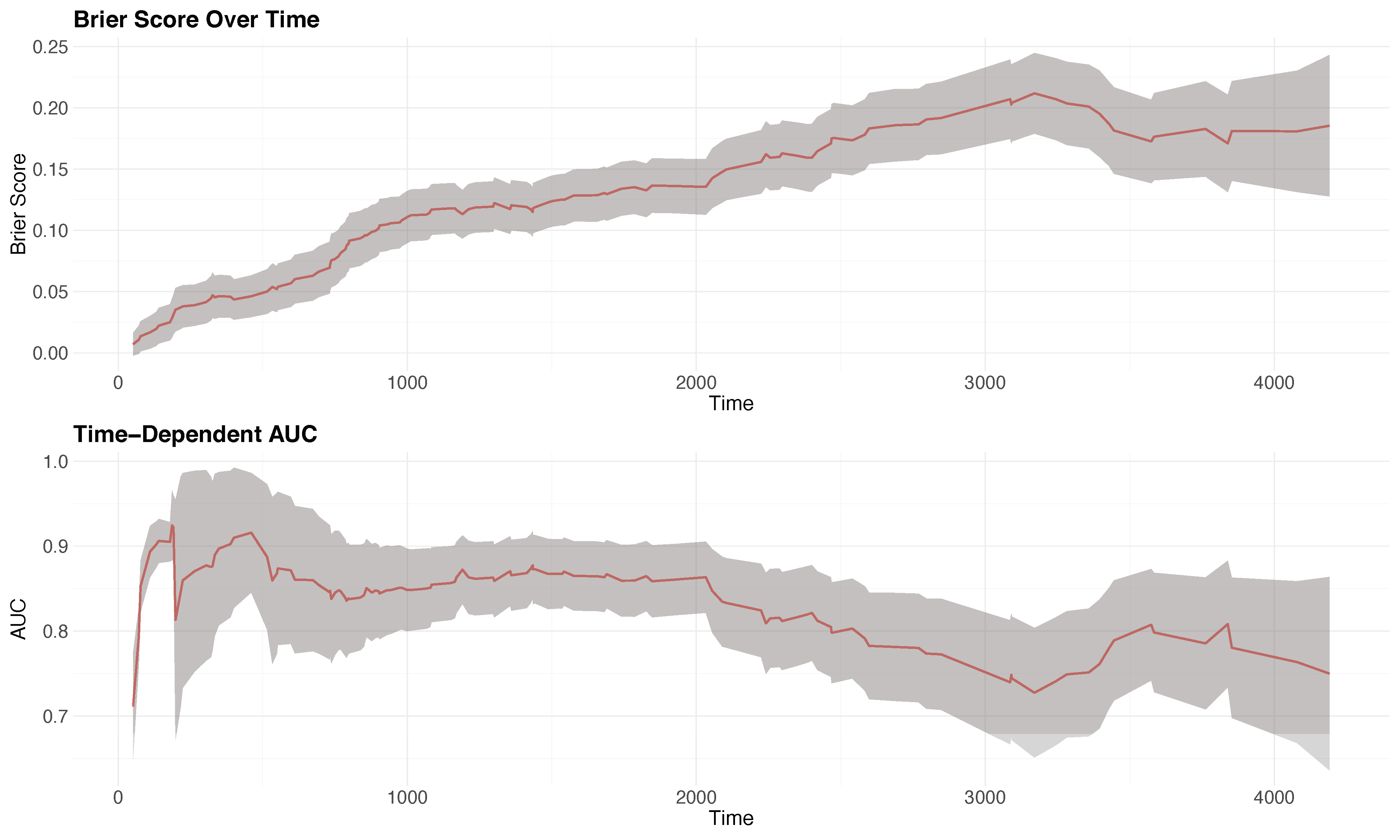
Tip:
Code generated using plot.brier.auc
Prediction
4 major ways prediction can be used
Prediction on test data:
predict(object, testdata)Prediction on “artificial data” for partial plots:
plot.variable,partialPrediction where a new data is substituted into the trained forest:
predict(object, testdata, outcome = "test")Restoring the forest:
predict(object, ...)
General call to predict
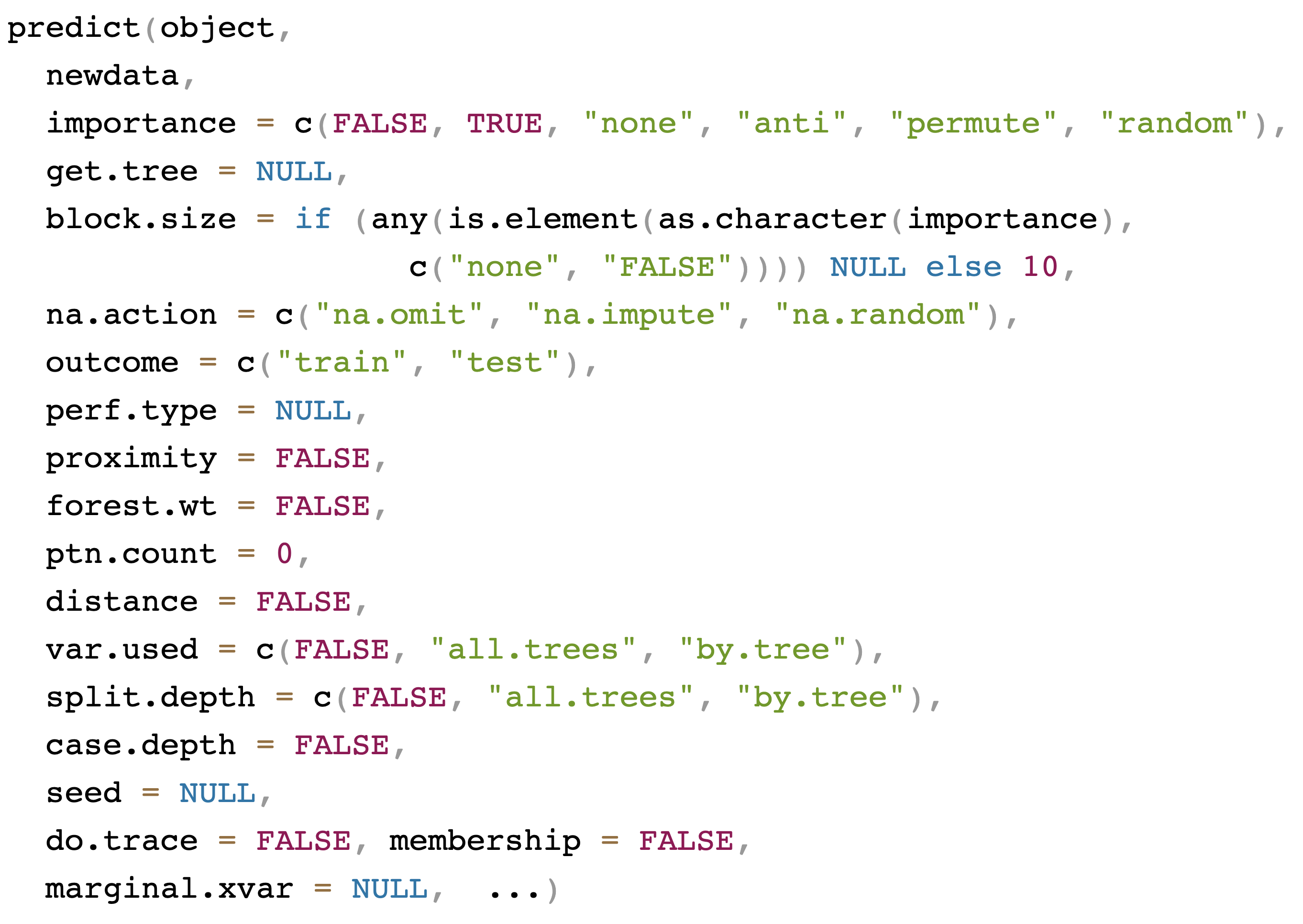
Prediction: canonical example
Veteran’s Administration Lung Cancer Trial
Randomized trial of two treatment regimens for lung cancer [2]
| trt | celltype | time | status | karno | diagtime | age | prior |
|---|---|---|---|---|---|---|---|
| 1 | 1 | 72 | 1 | 60 | 7 | 69 | 0 |
| 1 | 1 | 411 | 1 | 70 | 5 | 64 | 10 |
| 1 | 1 | 228 | 1 | 60 | 3 | 38 | 0 |
| 1 | 1 | 126 | 1 | 60 | 9 | 63 | 10 |
| 1 | 1 | 118 | 1 | 70 | 11 | 65 | 10 |
| 1 | 1 | 10 | 1 | 20 | 5 | 49 | 0 |
| 1 | 1 | 82 | 1 | 40 | 10 | 69 | 10 |
| 1 | 1 | 110 | 1 | 80 | 29 | 68 | 0 |
| 1 | 1 | 314 | 1 | 50 | 18 | 43 | 0 |
| 1 | 1 | 100 | 0 | 70 | 6 | 70 | 0 |
Prediction: canonical example
## veteran data (with factors)
data(veteran, package = "randomForestSRC")
veteran2 <- data.frame(lapply(veteran, factor))
veteran2$time <- veteran$time
veteran2$status <- veteran$status
## split the data into unbalanced train/test data (25/75)
## the train/test data have the same levels, but different labels
train <- sample(1:nrow(veteran2), round(nrow(veteran2) * .25))> summary(veteran2[train,])
trt celltype time status karno diagtime age prior
1:18 1: 8 Min. : 3.00 Min. :0.0000 60 :7 3 : 7 60 : 3 0 :27
2:16 2:10 1st Qu.: 29.25 1st Qu.:1.0000 40 :6 2 : 6 66 : 3 10: 7
3: 9 Median : 53.50 Median :1.0000 80 :6 5 : 3 67 : 3
4: 7 Mean :104.29 Mean :0.9412 70 :4 8 : 3 69 : 3
3rd Qu.:140.25 3rd Qu.:1.0000 90 :4 1 : 2 38 : 2
Max. :389.00 Max. :1.0000 30 :3 4 : 2 48 : 2
(Other):4 (Other):11 (Other):18 > summary(veteran2[-train,])
trt celltype time status karno diagtime age prior
1:51 1:27 Min. : 1.0 Min. :0.000 60 :20 4 :17 63 :10 0 :70
2:52 2:38 1st Qu.: 21.5 1st Qu.:1.000 70 :19 2 :13 62 : 8 10:33
3:18 Median : 83.0 Median :1.000 80 :18 3 :11 64 : 7
4:20 Mean :127.3 Mean :0.932 30 :11 5 :11 65 : 7
3rd Qu.:147.5 3rd Qu.:1.000 50 :11 11 : 5 68 : 5
Max. :999.0 Max. :1.000 40 :10 12 : 5 70 : 5
(Other):14 (Other):41 (Other):61 Prediction: canonical example
## train the forest and use this to predict on test data
o <- rfsrc(Surv(time, status) ~ ., veteran2[train, ])
pred <- predict(o, veteran2[-train , ])> print(o)
Sample size: 34
Number of deaths: 32
Number of trees: 500
Forest terminal node size: 15
Average no. of terminal nodes: 1
No. of variables tried at each split: 3
Total no. of variables: 6
Resampling used to grow trees: swor
Resample size used to grow trees: 21
Analysis: RSF
Family: surv
Splitting rule: logrank *random*
Number of random split points: 10
(OOB) CRPS: 57.73879973
(OOB) stand. CRPS: 0.14842879
(OOB) Requested performance error: 0.88246269> print(pred)
Sample size of test (predict) data: 103
Number of grow trees: 500
Average no. of grow terminal nodes: 1
Total no. of grow variables: 6
Resampling used to grow trees: swor
Resample size used to grow trees: 21
Analysis: RSF
Family: surv
CRPS: 54.3104615
stand. CRPS: 0.13961558
Requested performance error: 0.49749046Prediction: canonical example
## even harder ... factor level not previously encountered in training
veteran3 <- veteran2[1:3, ]
veteran3$celltype <- factor(c("newlevel", "1", "3"))
pred2 <- predict(o, veteran3)
print(pred2)
> Sample size of test (predict) data: 3
> Number of grow trees: 500
> Average no. of grow terminal nodes: 1
> Total no. of grow variables: 6
> Resampling used to grow trees: swor
> Resample size used to grow trees: 21
> Analysis: RSF
> Family: surv
> CRPS: 60.6330452
> stand. CRPS: 0.15586901
> Requested performance error: 0.5
## the unusual level is treated like a missing value but is not removed
print(pred2$xvar)
> trt celltype karno diagtime age prior
> 1 1 <NA> 60 7 69 0
> 2 1 1 70 5 64 10
> 3 1 3 60 3 38 0Restore
Useful feature of predict making it possible to restore the original training forest
After tuning the forest, you can use predict to obtain: \[
\begin{array}{ll}
\texttt{importance} & \text{Variable Importance (VIMP)} \\
\texttt{forest.wt} & \text{Construct your own estimator} \\
\texttt{proximity} & \text{RF proximity} \\
\texttt{distance} & \text{RF distance} \\
\texttt{get.tree} & \text{Plot a tree on your browser} \\
\texttt{var.used, split.depth} & \text{Statistics summarizing variable and tree splitting} \\
\end{array}
\]
Restore
Tip
predict() can restore an object from rfsrc() with a new specification
| y | cg23970089 | cg25176823 | cg24576735 | Chr19.20co.gain | TERT.promoter.status | SNP6 | U133a | grade | age | gender |
|---|---|---|---|---|---|---|---|---|---|---|
| Mesenchymal-like | 0.36518600 | 0.3244530 | 0.03255164 | No chr 19/20 gain | Mutant | Yes | Yes | G4 | 44 | female |
| Mesenchymal-like | 0.60246131 | 0.3626941 | 0.02215452 | No chr 19/20 gain | Mutant | Yes | Yes | G4 | 50 | male |
| Mesenchymal-like | 0.44011951 | 0.3679728 | 0.03369096 | No chr 19/20 gain | Mutant | Yes | No | G4 | 56 | female |
| Classic-like | 0.06834791 | 0.3974054 | 0.03623296 | No chr 19/20 gain | Mutant | Yes | Yes | G4 | 40 | female |
| Classic-like | 0.19843219 | 0.2736760 | 0.04060163 | No chr 19/20 gain | Mutant | Yes | Yes | G4 | 61 | female |
o <- rfsrc(y~., data = glioma)
o
Sample size: 878
Frequency of class labels: Classic-like=148, Codel=174, G-CIMP-high=249, G-CIMP-low=25, LGm6-GBM=41, Mesenchymal-like=215, PA-like=26
Number of trees: 500
Forest terminal node size: 1
Average no. of terminal nodes: 73.79
No. of variables tried at each split: 36
Total no. of variables: 1241
Resampling used to grow trees: swor
Resample size used to grow trees: 555
Analysis: RF-C
Family: class
Splitting rule: gini *random*
Number of random split points: 10
(OOB) Brier score: 0.02270041
(OOB) Normalized Brier score: 0.18538666
(OOB) AUC: 0.99380046
(OOB) Log-loss: 0.33499374
(OOB) Requested performance error: 0.06719818, 0.04054054, 0.01149425, 0.00803213, 0.52, 0.48780488, 0.05581395, 0.15384615
Confusion matrix:
predicted
observed Classic-like Codel G-CIMP-high G-CIMP-low LGm6-GBM Mesenchymal-like PA-like class.error
Classic-like 143 0 0 0 0 5 0 0.0338
Codel 0 172 2 0 0 0 0 0.0115
G-CIMP-high 0 2 247 0 0 0 0 0.0080
G-CIMP-low 0 0 13 12 0 0 0 0.5200
LGm6-GBM 0 0 1 0 21 19 0 0.4878
Mesenchymal-like 12 0 0 0 0 203 0 0.0558
PA-like 0 0 0 0 0 4 22 0.1538
(OOB) Misclassification rate: 0.06605923
Random-classifier baselines (uniform):
Brier: 0.12244898 Normalized Brier: 1 Log-loss: 1.94591015p <- predict(o, perf.type = "brier")
p
Sample size of test (predict) data: 878
Number of grow trees: 500
Average no. of grow terminal nodes: 73.79
Total no. of grow variables: 1241
Resampling used to grow trees: swor
Analysis: RF-C
Family: class
Brier score: 0.02270041
Normalized Brier score: 0.18538666
AUC: 0.99380046
Log-loss: 0.33499374
Requested performance error: 0.18538666, 0.03943965, 0.01378517, 0.02434763, 0.01397218, 0.02309749, 0.06058674, 0.0101578
Confusion matrix:
predicted
observed Classic-like Codel G-CIMP-high G-CIMP-low LGm6-GBM Mesenchymal-like PA-like class.error
Classic-like 143 0 0 0 0 5 0 0.0338
Codel 0 172 2 0 0 0 0 0.0115
G-CIMP-high 0 2 247 0 0 0 0 0.0080
G-CIMP-low 0 0 13 12 0 0 0 0.5200
LGm6-GBM 0 0 1 0 21 19 0 0.4878
Mesenchymal-like 12 0 0 0 0 203 0 0.0558
PA-like 0 0 0 0 0 4 22 0.1538
(OOB) Misclassification rate: 0.06605923
Random-classifier baselines (uniform):
Brier: 0.12244898 Normalized Brier: 1 Log-loss: 1.94591015 Restore
Create your own estimator for regression
## create your own estimator for regression
o <- rfsrc(mpg~.,mtcars)
fwt <- predict(o, forest.wt="oob")$forest.wt
yhat <- c(fwt %*% o$yvar)
## compare to the OOB ensemble
print(summary(yhat - o$predicted.oob))
Min. 1st Qu. Median Mean 3rd Qu. Max.
-3.908e-14 -1.643e-14 -1.066e-14 -1.060e-14 -3.553e-15 1.066e-14
## compare prediction error to OOB ensemble
print(o)
Sample size: 32
Number of trees: 500
Forest terminal node size: 5
Average no. of terminal nodes: 3.476
No. of variables tried at each split: 4
Total no. of variables: 10
Resampling used to grow trees: swor
Resample size used to grow trees: 20
Analysis: RF-R
Family: regr
Splitting rule: mse *random*
Number of random split points: 10
(OOB) R squared: 0.76585425
(OOB) Requested performance error: 8.50513415
print(mean((yhat - o$yvar)^2))
8.505134Partial plots
Partial plots are used to understand the effect of a feature on the outcome and are essentially prediction tools
Consider nonparametric regression \[ F(x_1,\ldots,x_p) = E[Y|(x_1,\ldots,x_p)] \] To study variable \(j\) we plot \(F_j(x)\) over \(x\) where \[ F_j(x) = \frac{1}{n}\sum_{i=1}^nF_{\!\text{ens}}(x_1=x_{i1},\ldots,\color{red}{x_j=x},\color{black}\ldots,x_p=x_{ip}) \] and \[ \Bigg\{\Big(x_1=x_{i1},\ldots,\color{red}{x_j=x}\color{black},\ldots,x_p=x_{ip}\Big):\hskip10pt i=1,\ldots,n\Bigg\} \] are “test” data values equal to the training data but where coordinate \(j\) is set to \(x\)
General call to plot.variable

General call to partial

Survival example: peakVO2
The data involve 2231 patients with systolic heart failure who underwent cardiopulmonary stress testing at the Cleveland Clinic. The primary end point was all-cause death. In total, 39 variables were measured for each patient, including baseline clinical values and exercise stress test results. A key variable of interest is peak VO2 (mL/kg per min), the peak respiratory exchange ratio [3]
| ttodead | died | bun | sodium | hgb | glucose | male | black | crcl | age | betablok | dilver | nifed | acei | angioten.II | anti.arrhy | anti.coag | aspirin | digoxin |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1.207392 | 0 | 25.00000 | 141.0000 | 14.60000 | 89.00000 | 1 | 0 | 49.92778 | 74 | 1 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 |
| 3.611225 | 0 | 20.00000 | 138.0000 | 13.10000 | 90.00000 | 0 | 0 | 60.35000 | 77 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 1 |
| 0.238193 | 1 | 29.68983 | 139.8368 | 12.87517 | 98.74435 | 1 | 0 | 33.61172 | 79 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| 10.513347 | 0 | 27.87734 | 139.4590 | 13.16225 | 136.87414 | 1 | 0 | 69.25114 | 67 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 | 1 |
| 3.712526 | 1 | 39.00000 | 143.0000 | 12.50000 | 104.00000 | 1 | 0 | 44.67685 | 79 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 0 |
Survival example: peakVO2
We can use plot.variable to visualize \(F_j(x)\)
o <- rfsrc(Surv(ttodead,died)~., peakVO2)
## partial effect for age
plot.variable(o, surv.type = "surv", xvar.names = "age",
time = 2, partial = TRUE)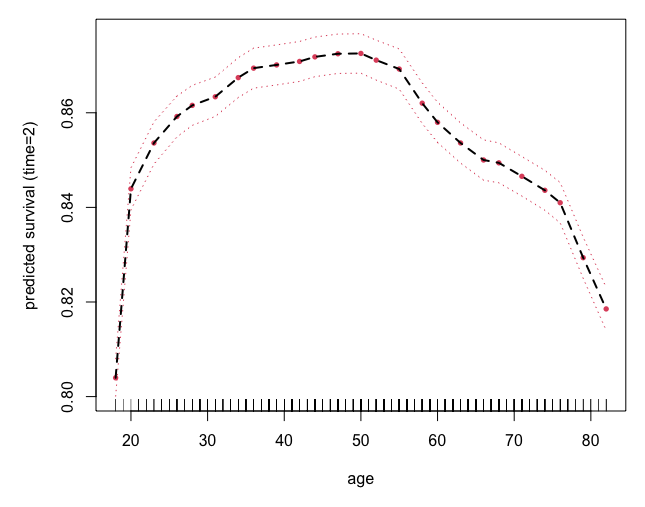
Survival example: peakVO2
## partial effect for age
plot.variable(o, surv.type = "surv", xvar.names = "age",
smooth.lines = TRUE,
time = 2, partial = TRUE)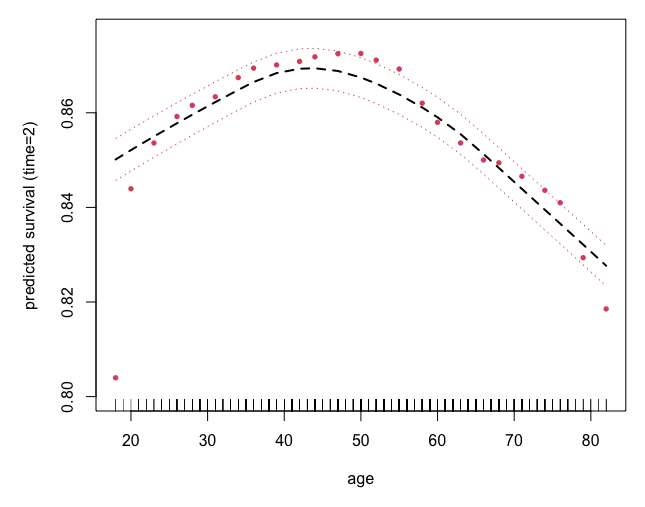
Survival example: peakVO2
## partial effect for age
plot.variable(o, surv.type = "rel.freq", xvar.names = "age",
smooth.lines = TRUE,
partial = TRUE)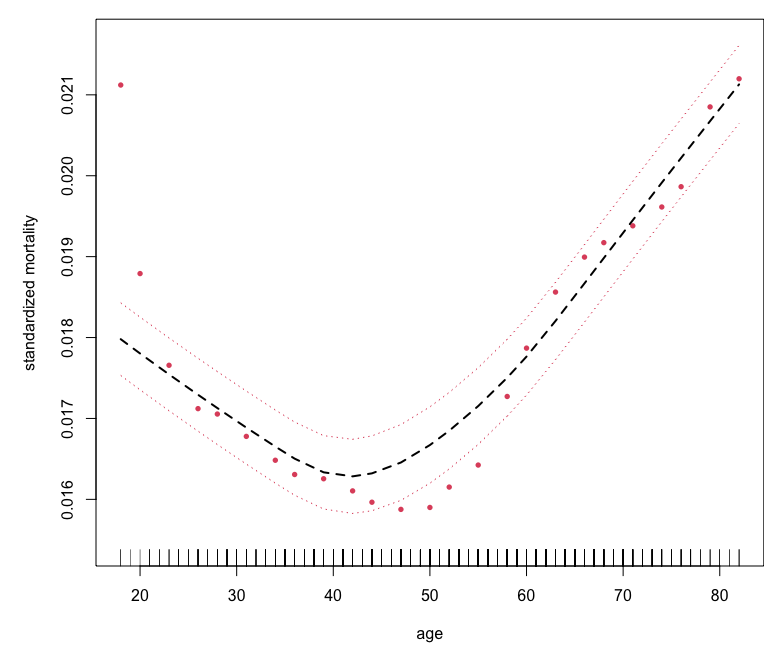
Survival example: peakVO2
We can use partial and get.partial.plot.data to obtain values of \(F_j(x)\)
## compare the two plots
par(mfrow=c(1,2))
plot(lowess(pdta.m$x, pdta.m$yhat, f = 2/3),
type = "l", xlab = "peak VO2", ylab = "adjusted mortality")
rug(o$xvar$peak.vo2)
matplot(pdta.s$partial.time, t(pdta.s$yhat), type = "l", lty = 1,
xlab = "years", ylab = "peak VO2 adjusted survival")
legend("bottomleft", legend = paste0("peak VO2 = ", pvo2),
bty = "n", cex = .75, fill = 1:5)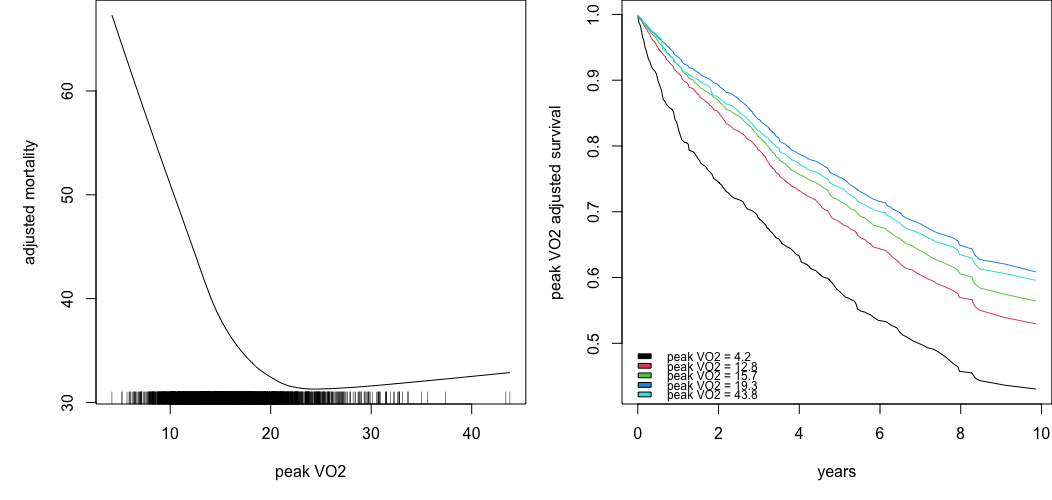
Outline
Part I: Training
- Quick start
- Data structures allowed
- Training (grow) with examples
(regression, classification, survival)
Part II: Inference and Prediction
- Inference (OOB)
- Prediction Error
- Prediction
- Restore
- Partial Plots
Part III: Variable Selection
- VIMP
- Subsampling (Confidence Intervals)
- Minimal Depth
- VarPro
Part IV: Advanced Examples
- Class Imbalanced Data
- Competing Risks
- Multivariate Forests
- Missing data imputation